MP252 AuthorsNeda Feizi Gilandeh1; Blessing Chisom Egbejiogu2; Kyle L. Wilhelm1; Kermit K. Murray2; Touradj Solouki1 Institutes1Baylor University, Waco, TX; 2Louisiana State University, Baton Rouge, LA IntroductionNative electrospray ionization mass spectrometry (n-ESI/MS) combined with ion mobility spectrometry (IMS) provides an opportunity to study indigenous forms of proteins in biological samples. Recently, we reported on combining n-ESI/MS …
Tag Archives: Ablation
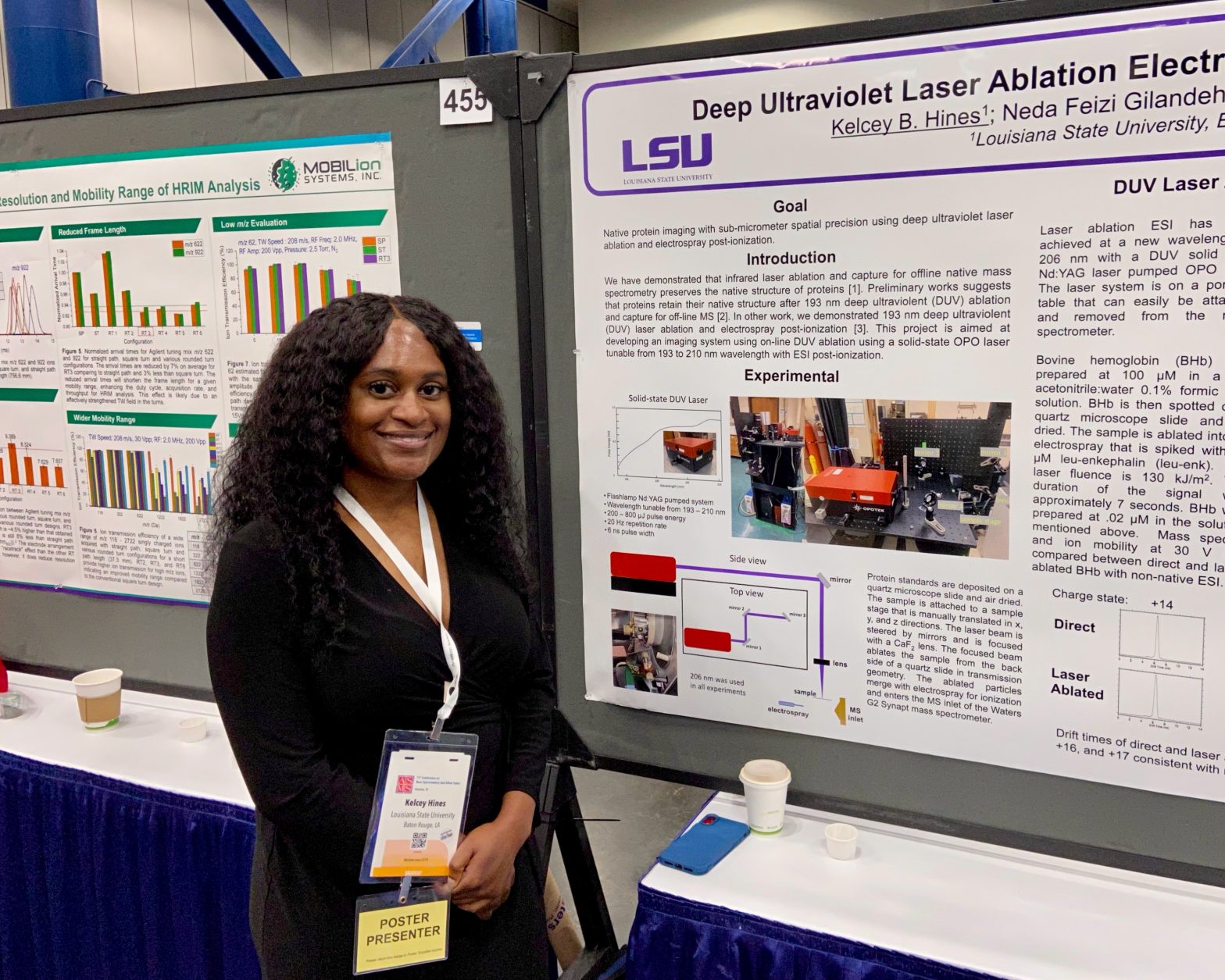
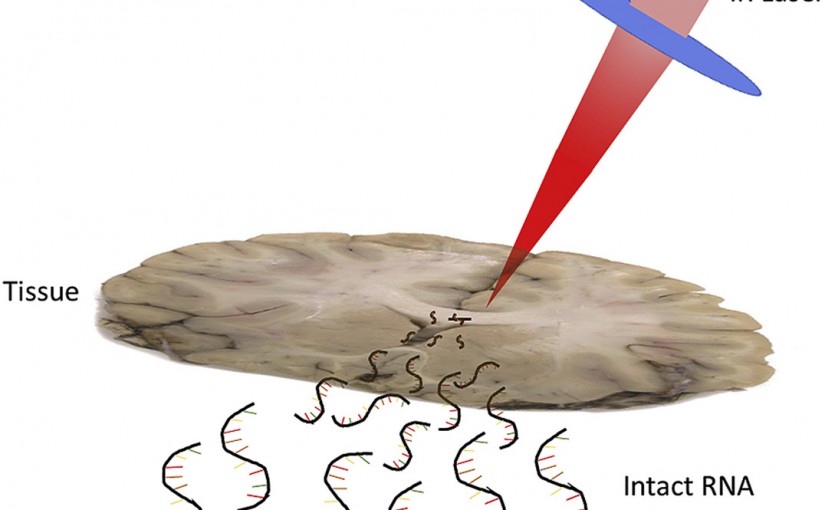
ASMS 2023: Deep Ultraviolet Laser Ablation Electrospray Ionization for Native Mass Spectrometry
WP 456 Murray Group ASMS 2023 Presentations
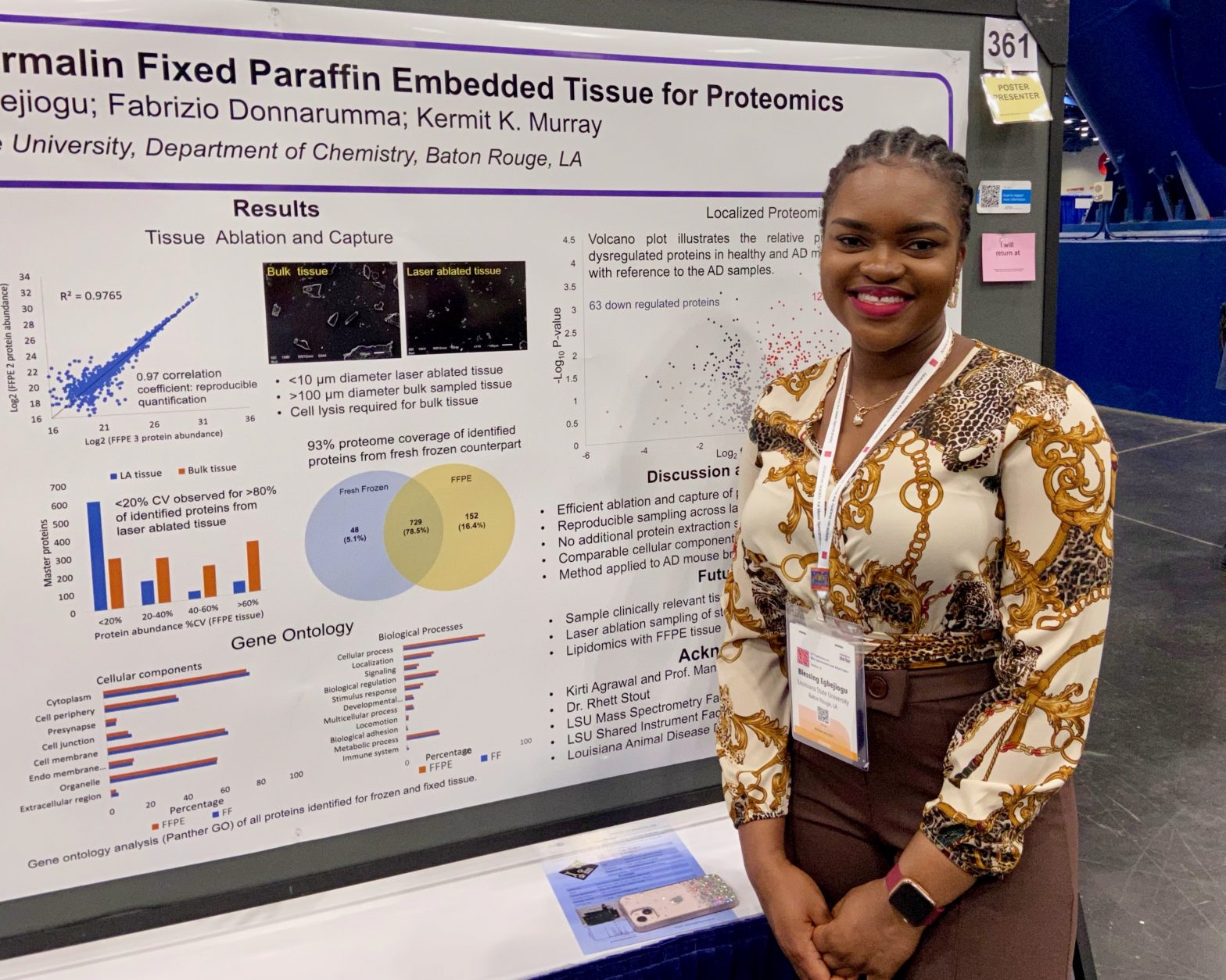
ASMS 2023: Localized Microsampling of Formalin Fixed Paraffin Embedded Tissue for Proteomics
TP361 Murray Group ASMS 2023 Presentations
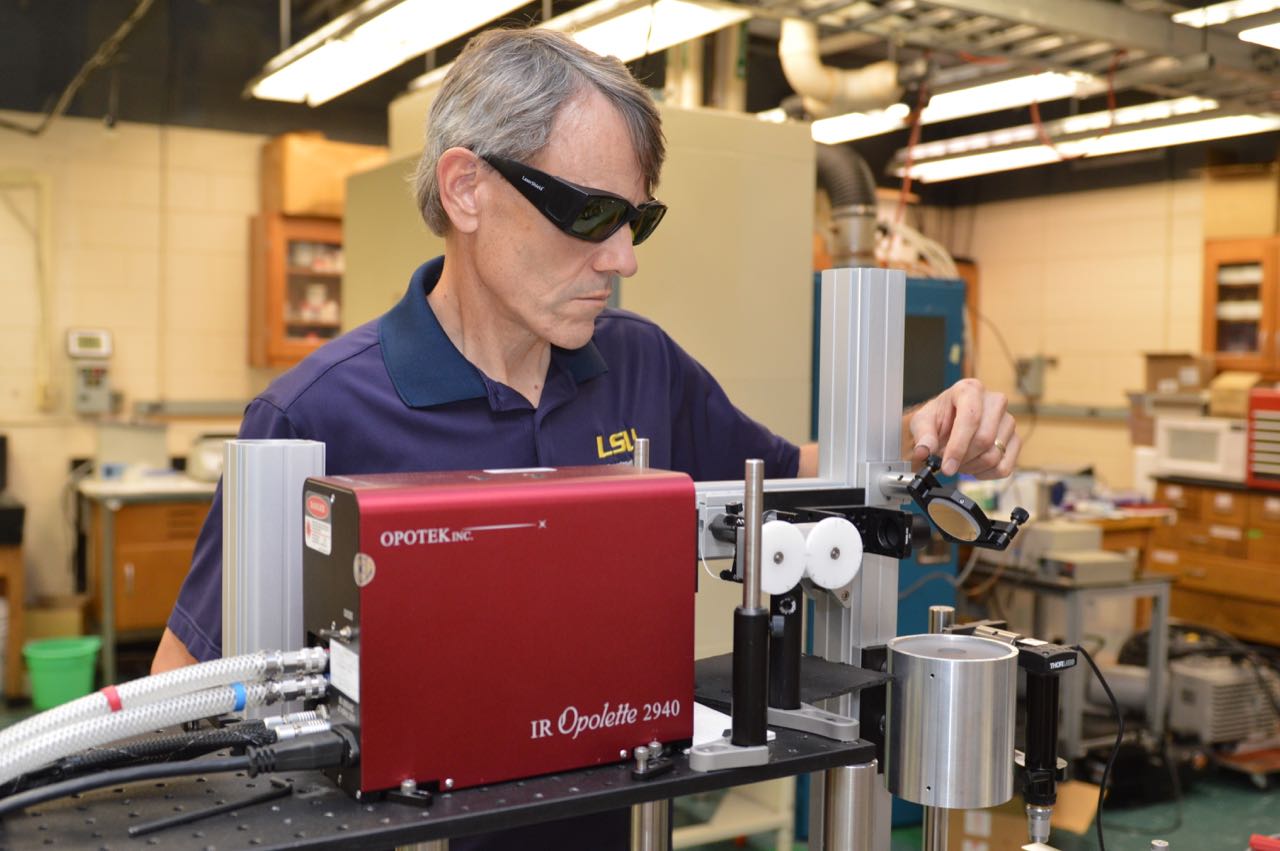
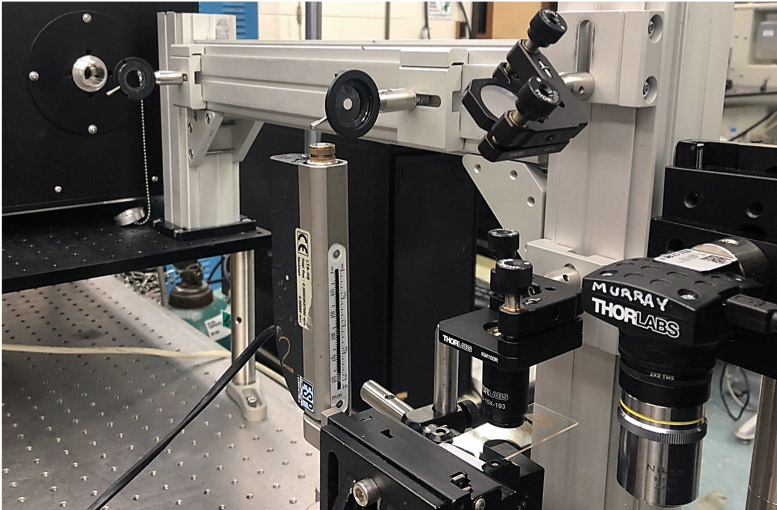
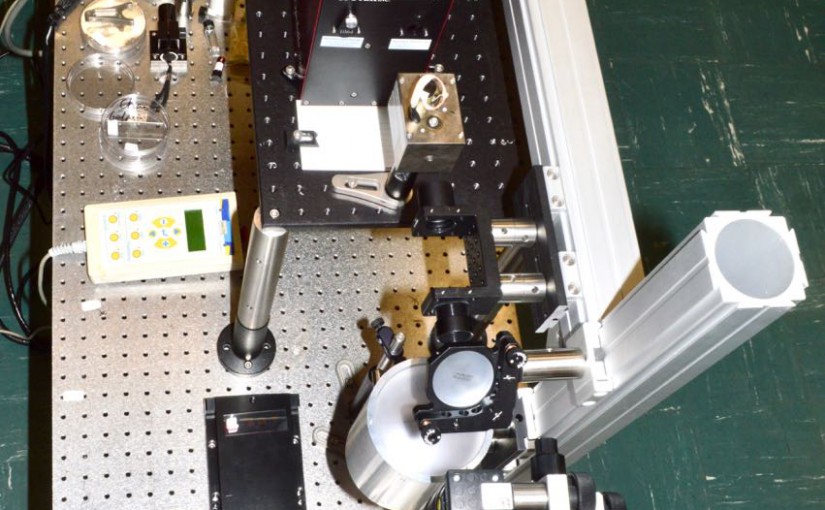
Opotek Customer Spotlight
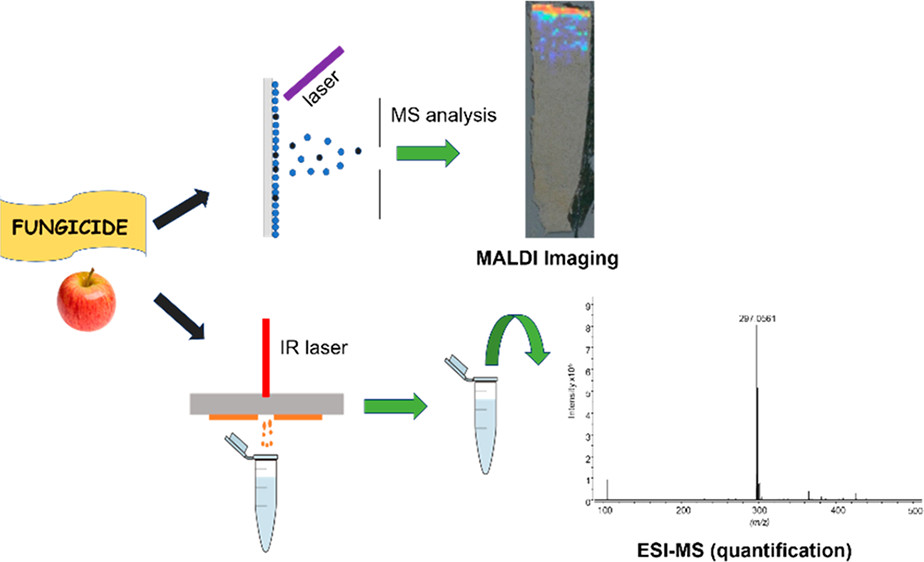
Matrix-Assisted Laser Desorption Ionization Imaging and Laser Ablation Sampling for Analysis of Fungicide Distribution in Apples
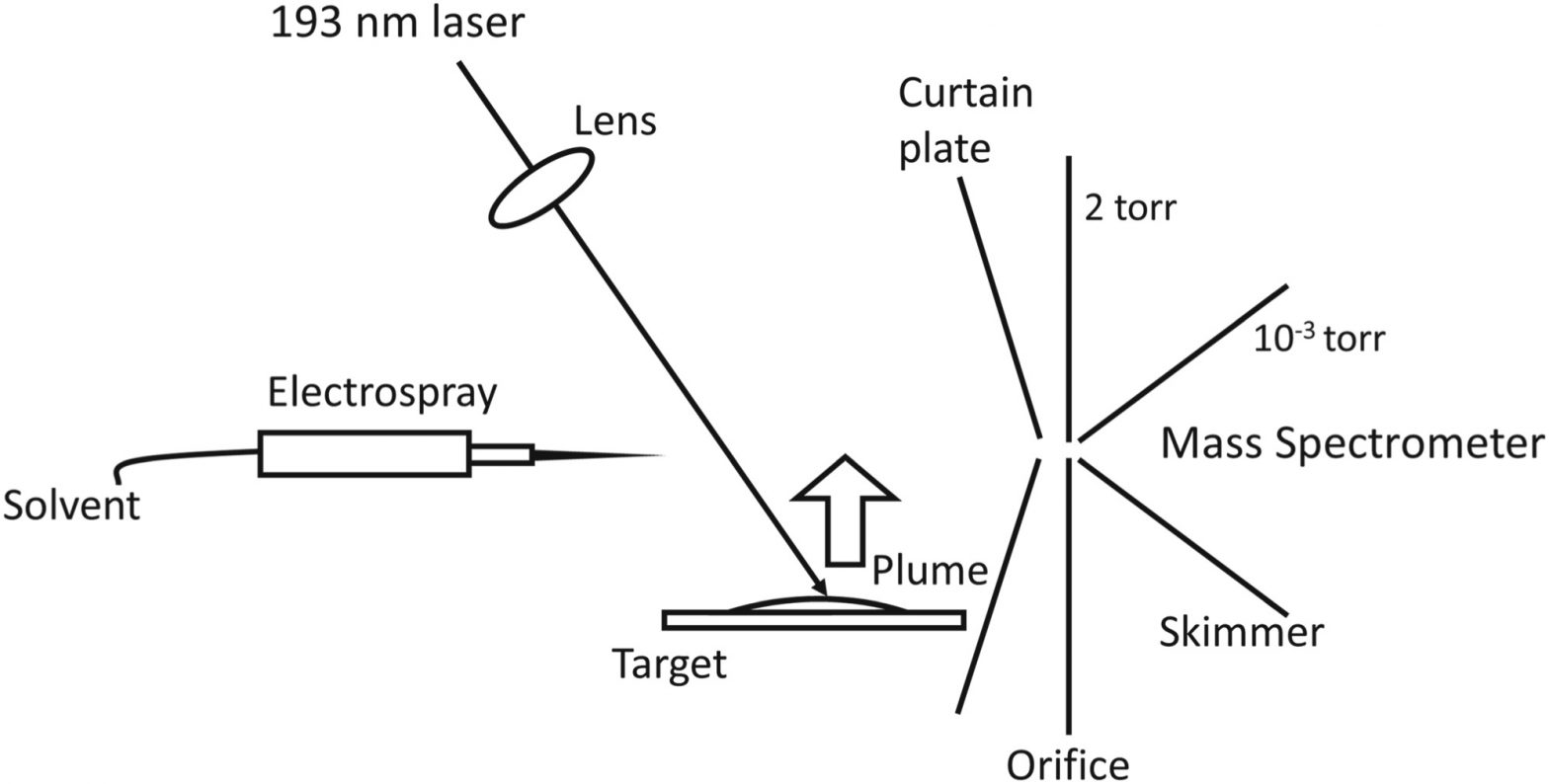
ASMS 2019: Deep-ultraviolet Laser Ablation Sampling for Mass Spectrometry
MOG 03:10pm