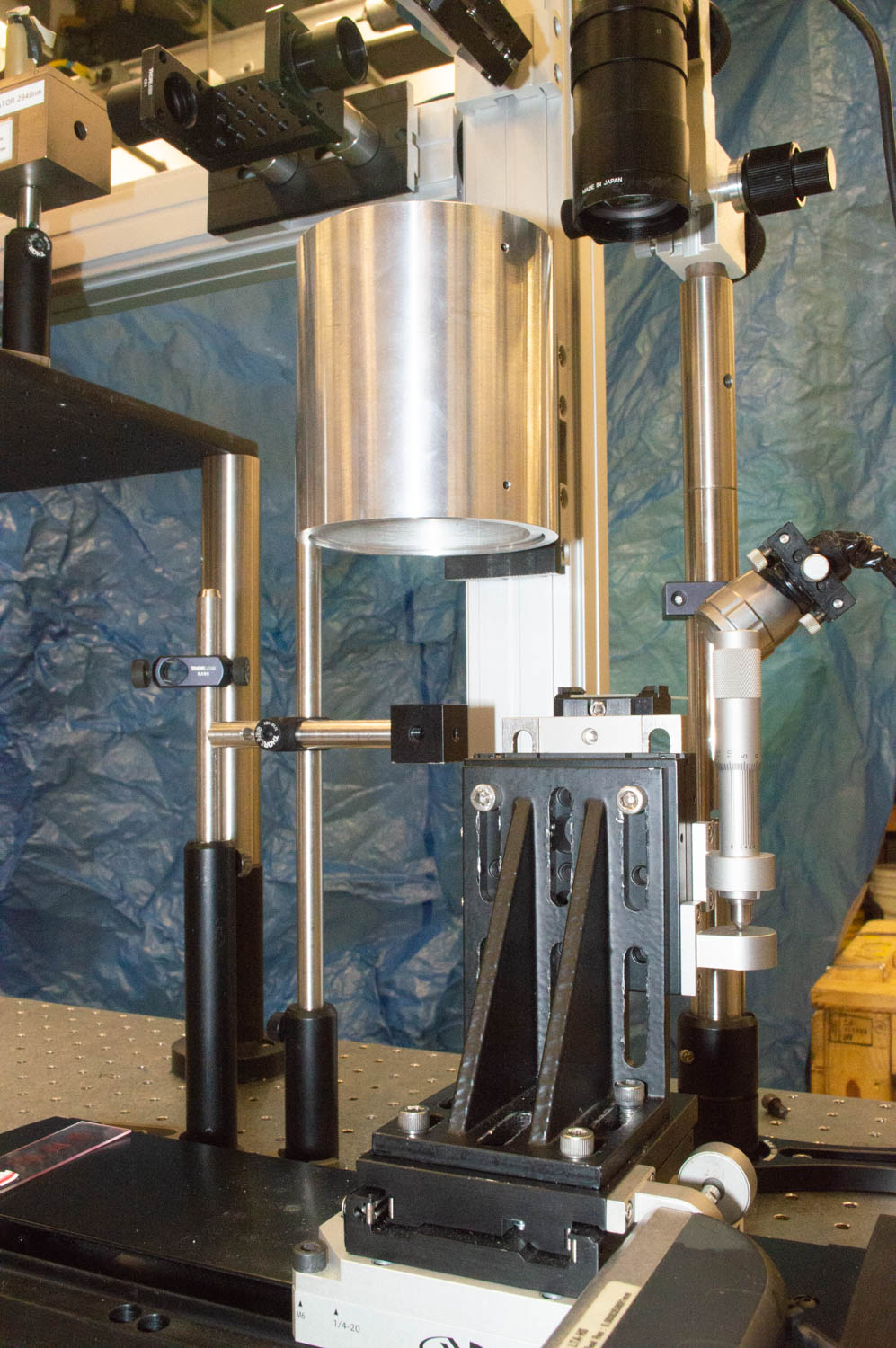
International Mass Spectrometry Conference, Florence, Italy, August 28, 2018 Louisiana State University: Kermit K. Murray, Fabrizio Donnarumma, Kelin Wang, Carson W. Szot, Chao Dong,Baylor University: Touradj Solouki, Michael E. Pettit Introduction Matrix-assisted laser desorption/ionization (MALDI) mass spectrometry imaging (MSI) is a powerful method for determining the location of biomolecules in tissue; however, protein identification and …
Continue reading “IMSC 2018: Infrared Laser Ablation Microsampling Coupled with MALDI Imaging”